Modeling
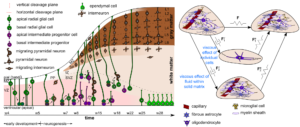
Accurate models that mathematically describe how tissue deformations are related to tissue stresses and how those relations change with time are a prerequisite to achieve useful in silico predictions. Due to its ultrasoft nature, brain tissue undergoes large deformations even under its own weight, and certainly during injury- or disease-relevant situations. Therefore, we develop models within the framework of finite deformation continuum mechanics. While one major drawback of previous models has been that they were based on poor, rather limited experimental data and could not capture the highly heterogeneous and dynamic nature of brain tissue, we develop mechanical models hand in hand with the versatile set of experiments. We aim to capture the complex and unique characteristics of this ultrasoft, heterogeneous, and active tissue through multi-scale and multi-physics approaches to couple tissue elasticity, viscosity, porosity, cell behavior, and volume growth through appropriate constitutive and evolution laws. Like this, we aim to bridge the scales from microstructural changes on the cellular scale to the mechanical response on the continuum scale.